1. The company OncoDNA provides test OncoDeep: 70 genes + predicting immunotherapy response.
https://www.oncodna.com/en/immunotherapy/
Price: 2990Eur.
Are there any studies predict immune therapy in other laboratories? Is this study useful?
3. Here (https://www.cegat.de/en/diagnostics/tumor-diagnostics/ovarian-cancer/) it is reported that "The detection of a somatic or germline mutation in the BRCA1 or BRCA2 genes is a requirement for treatment with Olaparib (LynparzaTM)"
There are also 4 testing options.
"Option 1: BRCA1 and BRCA2 analysis in tumor tissue only
In option 1, only mutations in the tumor are analyzed, no differentiation between germline and somatic mutations can be made. We do not recommend this option, but will perform it when it is explicitly requested by a patient."
Is it necessary to do such a study to determine the possibility of treatment with Olaparib or is it enough to study Oncodeep (by OncoDNA)? But in Oncodeep only mutations in the tumor are analyzed.
Option 1: BRCA1 and BRCA2 analysis in tumor tissue only
Option 4: Somatic Tumor Panel
https://www.oncodna.com/en/immunotherapy/
Price: 2990Eur.
Are there any studies predict immune therapy in other laboratories? Is this study useful?
2. By the way, in Moscow (Russia) a study of 50 genes costs $480. Can this research give some useful information? Or these 50 genes is very limited and such information is not sufficient?
3. Here (https://www.cegat.de/en/diagnostics/tumor-diagnostics/ovarian-cancer/) it is reported that "The detection of a somatic or germline mutation in the BRCA1 or BRCA2 genes is a requirement for treatment with Olaparib (LynparzaTM)"
There are also 4 testing options.
"Option 1: BRCA1 and BRCA2 analysis in tumor tissue only
In option 1, only mutations in the tumor are analyzed, no differentiation between germline and somatic mutations can be made. We do not recommend this option, but will perform it when it is explicitly requested by a patient."
Is it necessary to do such a study to determine the possibility of treatment with Olaparib or is it enough to study Oncodeep (by OncoDNA)? But in Oncodeep only mutations in the tumor are analyzed.
Option 1: BRCA1 and BRCA2 analysis in tumor tissue only
Option 4: Somatic Tumor Panel
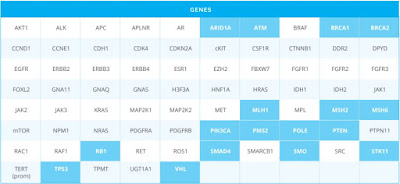





Both Foundation Medicine and Caris also offer the microsatellite instability and Tumor Mutational burden testing, as well as PD-L1 expression, and they sequence a larger number of genes, but they are also more expensive.
ReplyDeleteOne thing offered by OncoDNA that isn't being offered by these other companies is CD8 expression levels.
Most newly diagnosed tumors have low Tumor Mutational Burden, but in recurrent tumors treated with prior TMZ there is more of a chance of seeing high Tumor Mutational Burden (that is, hypermutation).
OncoDEEP could be a useful test. Advantages of OncoDEEP would be lower cost and a few tests (like CD8 expression) not offered by other companies.
On the downside, it sequences far fewer genes compared to the other mentioned companies. The 75 gene panel includes most of the commonly mutated genes seen in GBM, with the exception of NF1 and PIK3R1 (it doesn't include these).
2. Like OncoDEEP, this 50 gene panel includes the most commonly mutated GBM genes, except for NF1, PIK3R1, and the TERT promoter. At $480 this would be very cost effective, but you're not getting all the additional testing offered by OncoDNA and the other companies.
3. BRCA mutations (often germline) are common in ovarian cancer, but very rare in GBM. Ovarian cancer is associated with germline BRCA mutation, whereas GBM isn't. So the BRCA testing wouldn't be a high priority for GBM. PARP inhibitors like olaparib and rucaparib are only approved for BRCA-mutant ovarian cancer, so any use in GBM would be off-label (the vast majority of GBMs don't have a BRCA mutation). Trials currently underway will clarify whether PARP inhibiton is a useful strategy for GBM.
Stepnen,
Deletehow important is the testing of genes NF1, PIK3R1? As testing them forced to order a much more expensive test(
Hi Stephen,
ReplyDeleteIt's a nice summary of OncoDEEP test!
Hi All,
If you have any questions, feel free to contact me at m.schlogel @ oncodna.com
Hi, Matthieu!
DeleteIs it possible to add to OncoDeep testing of mutation EGFRvIII and genes NF1, PIK3R1? And how much will it total cost?
According to Brennan et al. 2013 https://www.ncbi.nlm.nih.gov/pubmed/24120142
ReplyDelete(Supplemental Table S2)
The 6 most common somatic mutations, mutated in at least 10% of GBM are:
TP53 (34%)
EGFR (33%)
PTEN (32%)
NF1 (14%)
PIK3CA (12%)
PIK3R1 (12%)
This was based on sequencing the exome of 291 GBMs. Additionally the TERT promoter is mutated in about 83% of GBMs.
NF1 inactivates RAS. When NF1 is inactivated by mutation, it leads to overactive RAS signaling and downstream MEK and ERK signaling. Key druggable targets in this case would be MEK (target of drugs like trametinib and cobimetinib) and mTOR complex 1 (target of drugs like everolimus, rapamycin, temsirolimus).
https://www.ncbi.nlm.nih.gov/pubmed/24913553
The problem is that none of these drugs are approved for GBM, and probably have difficulty crossing the blood-brain barrier very well.
PIK3CA and PIK3R1 mutations could both increase PI3K -> AKT -> mTOR signaling.
I might be inclined to try a rapamycin + hydroxy/chloroquine combination, even in the absence of any genetic testing.
Testing for NF1 mutation would mainly be important if there was any hope of getting access to a MEK inhibitor. Trametinib and cobimetinib are melanoma drugs, and for that indication they cost on the order of $7500 - $9000 (USD 2014) per month, so clearly a huge expense if paying out-of-pocket and for uncertain benefit because much of this type of drug gets stopped at the blood brain barrier.
https://www.ncbi.nlm.nih.gov/pubmed/24875464
The EGFRvIII mutation is a deletion of exons 2-7 of the EGFR gene, so that should be included in OncoDEEP which covers EGFR (but would be good to verify that).
Do you think this combination (rapamycin + hydroxy/chloroquine) is promising, even if the patient starts it after radiation therapy and with a large residual tumor after surgery? Because in studies patients began to take this combination from the first day of radiotherapy.
DeleteIt's hard to say for sure, but theoretically yes this combo could have activity in combination with chemotherapy or even on its own.
DeleteThere is a trial being planned in Taiwan to formally test the rapamycin + hydroxychloroquine combo.
https://clinicaltrials.gov/ct2/show/NCT03008148